In the marine environment, fatty acids are found with a variety of structures and many organisms have unique fatty acid profiles in their fat storage tissues. These profiles or signatures resemble the fatty acid profiles of the diets of the consumer. One aspect of our work in this area is currently focused on using these fatty acid profiles to recognize differences in feeding ecology in competing forage fish species in the Gulf of Alaska. We are also using controlled feeding studies to understand the ways in which consumers subtly modify the fatty acid profiles of their prey so that we can better estimate recent diets using fatty acid signatures. Last, in collaboration with Jeff Bromaglin at USGS, we are investigating methods to improve the quantitative fatty acid signature analysis (QFASA) model used to estimate predator diets.
More recently, we have begun to investigate the stable carbon isotopes of fatty acids as specific markers of diets, 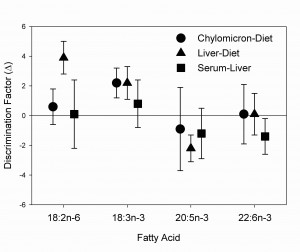 using compound-specific isotope analysis. While concentrations of fatty acids will change from prey to predator, changing the fatty acid profile, we expect very little modification in the isotopic signature of fatty acids, particularly essential fatty acids. Using Atlantic pollock as a model species, we are currently investigating the fractionation of dietary fatty acids during incorporation into fish tissues.
using compound-specific isotope analysis. While concentrations of fatty acids will change from prey to predator, changing the fatty acid profile, we expect very little modification in the isotopic signature of fatty acids, particularly essential fatty acids. Using Atlantic pollock as a model species, we are currently investigating the fractionation of dietary fatty acids during incorporation into fish tissues.
Recent Publications:
O’Donovan, S.A, Budge, S.M., Hobson, K.A., Kelly, A.P. and Derocher, A.E. 2018. Intrapopulation variability in wolf diet revealed using a combined stable isotope and fatty acid approach. Ecosphere 9: e02420. 10.1002/ecs2.2420.
Conners, M.G., Goetsch, C., Budge, S.M., Walker, W.A., Mitani, Y., Costa, D.P. and Shaffer, S.A. 2018. Fisheries exploitation by a marine predator quantified with lipid analysis. Frontiers in Marine Science. doi: 10.3389/fmars.2018.00113.
Bromaghin, J.F., Budge, S.M., Thiemann, G.W. and Rode, K.D. 2017. Simultaneous estimation of diet composition and calibration coefficients with fatty acid signature data. Ecology and Evolution 00:1–12. DOI: 10.1002/ece3.3179.
Oxtoby, L.E., Budge, S.M., Iken, K., O’Brien, D.M. and Wooller, M.J. 2016. Feeding ecologies of key bivalve and polychaete species in the Bering Sea as elucidated by fatty acid and compound specific stable isotope analyses. Marine Ecology Progress Series 557: 161-175. https:/doi.org/10.3354/meps11863.
Budge, S.M., Aucoin, L.R., Ziegler, S.E. and Lall, S.P. 2016. Fractionation of stable carbon isotopes of tissue fatty acids in Atlantic pollock (Pollachius virens). Ecosphere 7: 1-16. DOI: 10.1002/ecs2.1437.
Bromaghin, J.F., Rode, K.D., Budge, S.M. and Thiemann, G.W. 2015. Distance measures and optimization spaces in quantitative fatty acid signature analysis. Ecology and Evolution 5: 1249-1262. DOI: 10.1002/ece3.1429
Wang, S.W., Budge, S.M., Iken, K., Gradinger, R.R., Springer, A.M. and Wooller, M.J. 2014. Importance of sympagic production to Bering Sea zooplankton as revealed from fatty acid-carbon stable isotope analyses. Marine Ecology Progress Series. doi: 10.3354/meps11076.
